Abstract
-
OBJECTIVES
- To estimate time-variant reproductive number (Rt) of coronavirus disease 19 based on either number of daily confirmed cases or their onset date to monitor effectiveness of quarantine policies.
-
METHODS
- Using number of daily confirmed cases from January 23, 2020 to March 22, 2020 and their symptom onset date from the official website of the Seoul Metropolitan Government and the district office, we calculated Rt using program R’s package “EpiEstim”. For asymptomatic cases, their symptom onset date was considered as -2, -1, 0, +1, and +2 days of confirmed date.
-
RESULTS
- Based on the information of 313 confirmed cases, the epidemic curve was shaped like ‘propagated epidemic curve’. The daily Rt based on Rt_c peaked to 2.6 on February 20, 2020, then showed decreased trend and became <1.0 from March 3, 2020. Comparing both Rt from Rt_c and from the number of daily onset cases, we found that the pattern of changes was similar, although the variation of Rt was greater when using Rt_c. When we changed assumed onset date for asymptotic cases (-2 days to +2 days of the confirmed date), the results were comparable.
-
CONCLUSIONS
- Rt can be estimated based on Rt_c which is available from daily report of the Korea Centers for Disease Control and Prevention. Estimation of Rt would be useful to continuously monitor the effectiveness of the quarantine policy at the city and province levels.
-
Keywords: COVID-19, Communicable disease, Epidemics, Seoul
INTRODUCTION
- As of March 27, 2020, there were 504,806 confirmed cases of a coronavirus disease 2019 (COVID-19) reported worldwide and 134 countries were reporting community transmission [1]. In Korea, a total of 9,332 confirmed cases were reported from January 20, 2020, when the first case was confirmed, to March 27, 2020. The daily number of confirmed cases in Korea increased rapidly after a large-scale cluster of COVID-19 cases occurred in mid-February at the Sincheonji Church in Daegu, reaching 909 newly confirmed cases per day on February 29, 2020. After February 29, 2020 the number of new cases has decreased, but small and large outbreaks are still being reported nationwide, and the total number of new cases outside of Daegu has increased as the immigration of COVID-19 from foreign countries increases [1].
- The reproduction number (R) is defined as the average number of infected people generated during the infectious period of an infected patient. It is an index to quantify the transmissibility of infectious disease. The R value varies over time during epidemic periods owing to various factors such as infection control strategies, pathogen characteristics, population immunity, and changes in contact behaviors between infectious and susceptible individuals. This time-varying R value is known as an instantaneous reproduction number or time-variant reproductive number (Rt). Therefore, observing changes in Rt is an important indicator for evaluating the effectiveness of infection control strategies and monitoring the spread of infection [2]. This study estimated Rt of COVID-19 using the information on confirmed cases in Seoul, Korea. It also summarized the results using the existing Rt statistical software package, EpiEstim.
METHODS
- Daily numbers of confirmed cases were obtained from COVID-19 status reports provided by the official website of Seoul city [3]. Moreover, information such as the presence or absence of symptoms and time of symptom onsets in the confirmed cases was collected from the official websites of Seoul district offices. A total of 329 cases were confirmed as infected from January 23, 2020, when the first case was confirmed in Seoul, to March 22, 2020. Table 1 shows the basic characteristics of these confirmed cases.
- Software packages such as R0 and EpiEstim that are optimized for estimating Rt have been developed [4,5]. In this study, EpiEstim was used because it was developed recently and it requires less computing time than the other [5]. Generation time, which is required to calculate Rt, is a time interval between infector’s infected date and its consecutive infectee’s infected date. However, generation time is usually estimated based on the difference between the time of symptom onset of the infector and the infectee, which is called serial interval, as it is often difficult to know the exact time of infection [6]. This study assumed the serial interval as gamma distributed with a mean of 3.96 days and a standard deviation of 4.75 days, which was reported in China [7].
- Rt was estimated based on the daily number of confirmed cases (Rt_c) and symptom onset (Rt_s) because it is difficult to identify the exact time of infection. Twelve confirmed cases before February 16, 2020 were excluded from the analysis because from January 23, 2020 to February 16, 2020, there was no Rt_c for a considerable period, thus we could not assume that they were infected from previous case within Seoul. Four more confirmed cases were excluded with symptoms but their onset dates of symptoms were missing. Finally, 313 confirmed cases were included in the analysis. In asymptomatic cases, the time of symptom onset was assumed to be the same as the time of diagnosis (n= 70). In sensitivity analysis, Rt was calculated by assuming the times of symptom onset in asymptomatic cases as confirmed date (TD)-2 days, TD-1 day, TD+1 day, and TD+2 days. Then, those calculated Rt were compared to the calculated Rt with assumption of asymptomatic cases’ symptom onset date as TD. The data analysis was performed using the R version 3.6.3 (https://cran.r-project.org/bin/windows/base/old/3.6.3/) and EpiEstim, and the median Rt and 95% confidence intervals (CIs) were obtained.
- Ethics statement
- This study uses data from official websites which are opened to public. So, this study is subject to institutional review board exception in accordance with article 13 of the enforcement ordinance (study using existing data or documents on subjects, etc.).
RESULTS
-
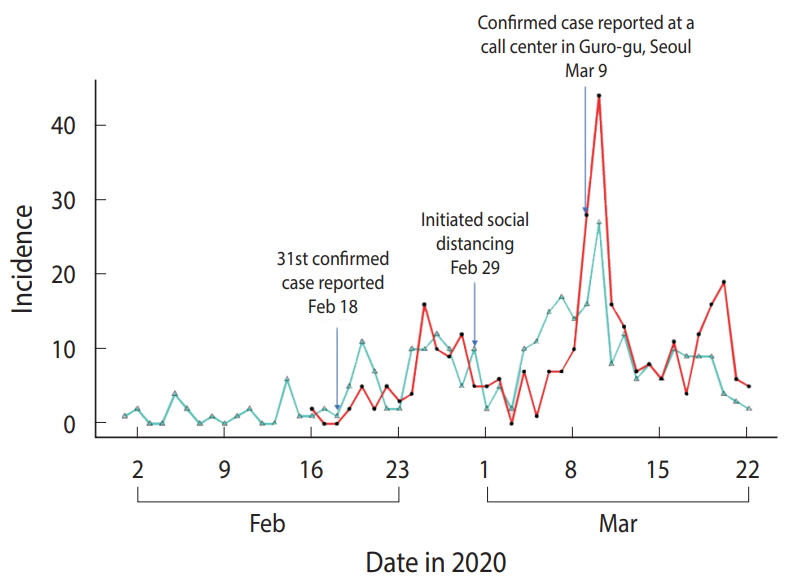
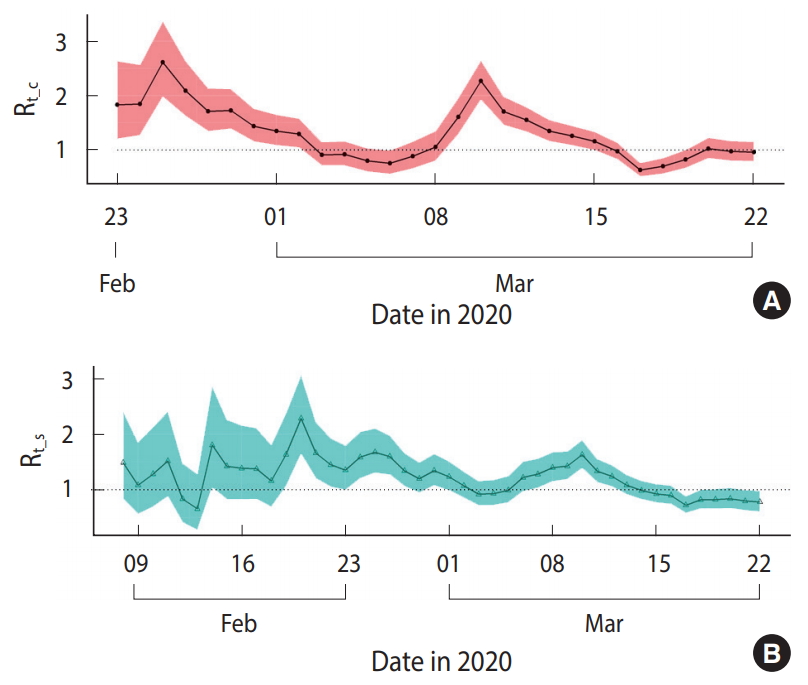
Figure 1 shows the distribution of the Rt_c and Rt_s in 313 confirmed cases included in the analysis. Figure 2A presents the median and 95% CI of Rt estimated using TD information. Assuming this Rt as Rt_c, Rt_c exhibited a decreasing trend from February 25, 2020 to March 6, 2020, which fell below 1 and then increased. It decreased again on March 10, 2020 and has shown a value of ≤ 1 after March 16, 2020. The median value and 95% CI of Rt, which was estimated using the Rt_s, are shown in Figure 2B. Assuming this Rt as Rt_s, Rt_s decreased to < 1 from February 20, 2020 to March 4, 2020, it increased shortly, and then decreased again from March 10, 2020 remaining below 1 after March 14, 2020.
-
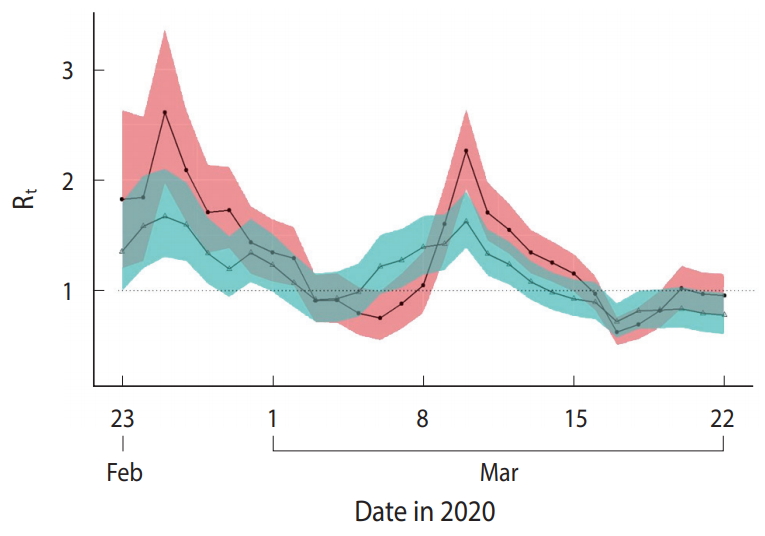
Figure 3 compares the mean Rt calculated based on the Rt_c and Rt_s. Overall, Rt_c and Rt_s showed similar changes in pattern; however, Rt_c showed more abrupt changes than Rt_s, and its highest and lowest values were estimated to be higher or lower than Rt_s, respectively. Moreover, the points in time when the uptrend or downtrend changes occurred were approximately 1 day after Rt_s, reflecting the lag time for testing and to confirm the diagnosis after symptom onset.
- In the primary analysis, the symptom onsets of asymptomatic patients (n= 70) were assumed to be the same as TD. To perform sensitivity analysis, Rt were calculated and compared, assuming the times of symptom onset were the same as TD-2 days, TD-1 day, TD+1 day, and TD+2 days, relative to TD. Under each assumption, the 95% CI of the Rt values overlapped considerably, and the changes over time showed a similar pattern, as shown in the Supplementary Material 1.
DISCUSSION
- In this study, the daily Rt of COVID-19 in Seoul from February 23, 2020 to March 22, 2020, was estimated using the data on the Rt_c and Rt_s. Daily Rt refers to the infectivity of newly confirmed cases on day t. In other words, an increase in Rt indicates that infected individuals are likely to transmit the disease more actively than previously, and this requires more intensive interventions for infection control. In contrast, current strategies to control infection are effective when Rt decreases. If the Rt constantly remains below 1, the epidemic will be disappeared [3].
- When the Rt was estimated using the Rt_c and Rt_s in Seoul, Rt values decreased from late February to early March and remained below 1 with some variations. This is possibly owing to the raised awareness of the public, enhanced infection control strategies, and social distancing, following a cluster outbreak that occurred at the Shincheonji Church in Daegu, Korea. The number of confirmed cases increased rapidly after another cluster outbreak was reported on March 10, 2020 at a call center in Guro-gu, Seoul. The Rt values remained higher than 1, reflecting the high transmission of the disease. However, the Rt decreased without further increase and has remained below 1 since March 10, 2020, after implementing infection control guidelines in high-risk workplaces, recommending the prohibition of mass gatherings, and limiting religious and public facility use as well as enhanced social distancing.
- The Rt values obtained using the Rt_c and Rt_s were comparable, despite of slight differences. When the Rt_c was used in the calculation, variations in the Rt were greater as the total number of confirmed cases increased rapidly. This was due to an increase in diagnostic testing after more investigations were completed following outbreaks or policy implementations. However, the variations in Rt declined when the values were calculated using the daily number of symptom onsets because symptom onsets are distributed widely before and after TD. Nevertheless, trends in Rt value changes were similar in both cases. In COVID-19, most patients developed mild initial symptoms, which made it difficult to determine the exact time of symptom onset. Therefore, the similar pattern of the estimated Rt indicate that the Rt using the Rt_c may be useful to estimate the pattern of infection transmission and to evaluate the effectiveness of the infection control strategies in the early stages.
- In the sensitivity analysis conducted in the asymptomatic cases with an assumption that the time of symptom onset was the same as TD-2 days to TD+2 days, the trend in estimated Rt showed similar changes in original data. The original data assumed that asymptomatic cases’ symptom onset dates were same as TD. Accordingly, the estimated Rt fells within the 95% CI of the original data. This indicates that assuming the time of symptom onset in asymptomatic cases as same as the date of confirmation is not significantly different from other assumptions in asymptomatic cases.
- There are several points to consider when interpreting Rt obtained in our study. First, the Rt may be underestimated as it gets closer to the latest date. The reason is that fewer values of symptom onset are used for the Rt value estimation than the actual number of confirmed cases due to lag time between symptom onset and confirmed date. Second, our study used Chinese serial interval as the generation time to estimate Rt. Since the generation time may be different from China, it is necessary to estimate Rt based on the estimated generation time in Korea, especially in Seoul. Third, the foreign immigration of confirmed cases was not considered. More accurate Rt prediction is possible if the infection transmission is traced accurately and confirmed whether there was an immigration of confirmed cases form outside of Seoul.
- In conclusion, the Rt estimated from using the Rt_c and Rt_s in Seoul were both useful in evaluating effectiveness of the infection control strategies. The values have remained below 1 since March 15, 2020, indicating a decreased rate of infection transmission from confirmed cases in the community. However, further studies should be conducted as the influx of confirmed cases from abroad and from other regions in Korea are increasing. Furthermore, the effectiveness of the infection control strategies should be monitored constantly at local and national levels, using Rt estimated with the Rt_c and Rt_s.
SUPPLEMENTARY MATERIALS
NOTES
-
CONFLICT OF INTEREST
The authors have no conflicts of interest to declare for this study.
-
FUNDING
This project was supported by a Government-wide R&D Fund Project for Infectious Disease Research (GFID), Republic of Korea (grant No. HG18C0088).
-
AUTHOR CONTRIBUTIONS
Conceptualization: BP, SGM. Data curation: YKK, SGM, BJN. Funding acquisition: BYC. Methodology: BP, JC, WSS, JHK. Writing – original draft: SGM, YKK, WSS, JHK, JC, BJN, BP, BYC. Writing – review & editing: SGM, YKK, WSS, JHK, JC, BJN, BP, BYC.
ACKNOWLEDGEMENTS
We wish to thank all the individuals who are struggling in the field of healthcare to overcome the COVID-19 outbreak. This study was performed under a research project named the Research and Development on Integrated Surveillance System for Early Warning of Infectious Diseases (RISEWIDs). The investigators of this project were Jaiyong Kim (Yonsei University), Sunworl Kim (National Medical Center), Eui Jung Kwon (National Health Insurance Review and Assessment Service), Dong Wook Kim (National Health Insurance Service), Moran Ki (National Cancer Center), Hyunjin Son (Pusan National University Hospital), Jong-Hun Kim (Sungkyunkwan University), Jin Yong Kim (Incheon Medical Center), Heeyoung Lee (Seoul National University Bundang Hospital), Boyoung Park (Hanyang University), Woo-Sik Son (National Institute for Mathematical Sciences), Jungsoon Choi (Hanyang University), Sunhwa Choi (National Cancer Center), Okyu Kwon (National Institute for Mathematical Sciences), Hyojung Lee (National Institute for Mathematical Sciences), Jong-Hoon Kim (International Vaccine Institute), Heecheon Kim (MISO Info Tech), and Bo Youl Choi (Hanyang University).
Figure 1.Distribution of the number of daily confirmed cases and symptom onsets in 313 confirmed coronavirus disease 2019 (COVID-19) cases in Seoul, Korea. The red line indicates the confirmed date, and the turquoise line indicates the onset date.
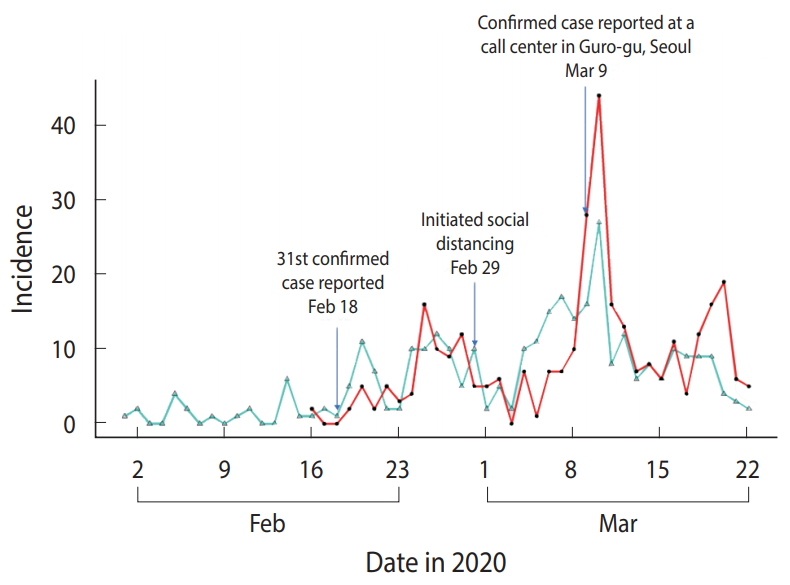
Figure 2.Reproductive number (Rt) with 95% percentile confidence intervals in Seoul from February 23, 2020 to March 22, 2020. (A) Based on the daily number of confirmed cases (Rt_c). (B) Based on the daily number of symptom onsets (Rt_s). The Rt_s begins on February 8, 2020.
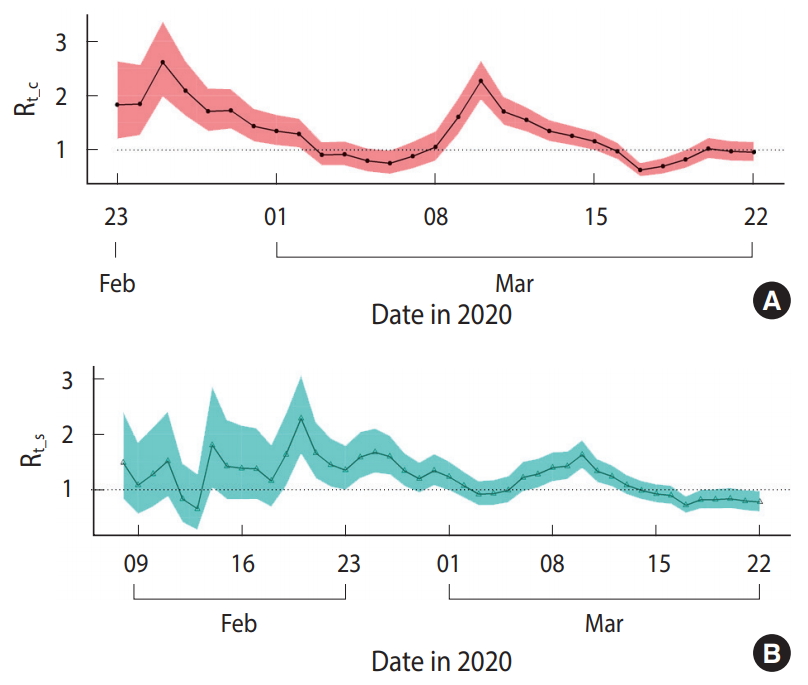
Figure 3.Comparison of the reproductive number (Rt) between the results based on daily number of confirmed cases (Rt_c) and results based on daily number of symptom onset (Rt_s) in Seoul, Korea from February 23, 2020 March 22, 2020.
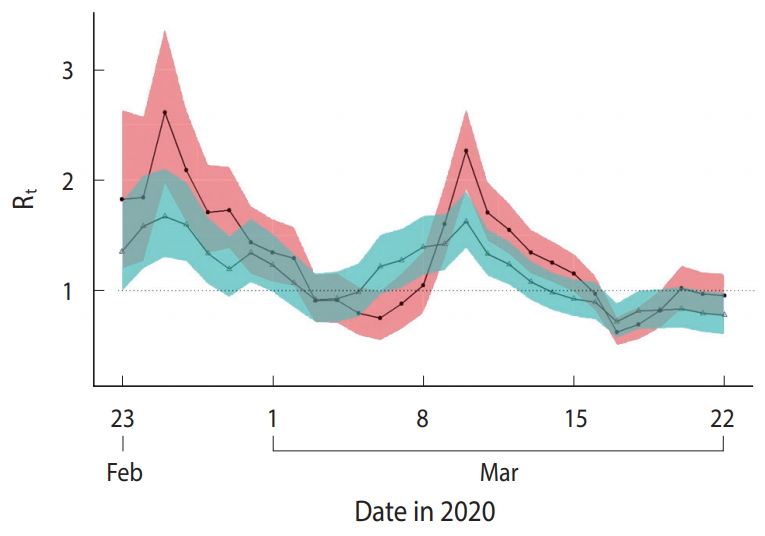
Table 1.Baseline characteristics of 329 confirmed coronavirus disease 2019 (COVID-19) case in Seoul from January 23, 2020 to March 22, 2020
|
Characteristics |
n (%) |
|
Sex |
|
|
Male |
161 (48.9) |
|
Female |
168 (51.1) |
|
Age (yr) |
|
|
<10 |
5 (1.5) |
|
10-19 |
11 (3.3) |
|
20-29 |
83 (25.2) |
|
30-39 |
51 (15.5) |
|
40-49 |
55 (16.7) |
|
50-59 |
70 (21.3) |
|
60-69 |
31 (9.4) |
|
70-79 |
15 (4.6) |
|
80-89 |
6 (1.8) |
|
≥90 |
2 (0.6) |
|
Days since symptoms to diagnosis (d)1
|
|
|
Asymptomatic |
70 (21.5) |
|
≤ 3 |
113 (34.8) |
|
4-7 |
85 (26.1) |
|
> 7 |
57 (17.5) |
REFERENCES
- 1. Korea Centers for Disease Control and Prevention. Coronavirus disease 19, Republic of Korea, cases in Korea; 2020 [cited 2020 Mar 27]. Available from: http://ncov.mohw.go.kr/ (Korean).
- 2. Lipsitch M, Bergstrom CT. Invited commentary: real-time tracking of control measures for emerging infections. Am J Epidemiol 2004;160:517-519.ArticlePubMedPDF
- 3. Seoul Metropolitan Government. Seoul coronavirus disease 19 (COVID-19), confirmer route; 2020 [cited 2020 Mar 27]. Available from: http://www.seoul.go.kr/coronaV/coronaStatus.do (Korean).
- 4. Obadia T, Haneef R, Boëlle PY. The R0 package: a toolbox to estimate reproduction numbers for epidemic outbreaks. BMC Med Inform Decis Mak 2012;12:147.ArticlePubMedPMCPDF
- 5. Cori A, Ferguson NM, Fraser C, Cauchemez S. A new framework and software to estimate time-varying reproduction numbers during epidemics. Am J Epidemiol 2013;178:1505-1512.ArticlePubMedPMCPDF
- 6. Vink MA, Bootsma MC, Wallinga J. Serial intervals of respiratory infectious diseases: a systematic review and analysis. Am J Epidemiol 2014;180:865-875.ArticlePubMedPDF
- 7. Du Z, Xu X, Wu Y, Wang L, Cowling BJ, Meyers LA. Serial interval of COVID-19 among publicly reported confirmed cases. Emerg Infect Dis 2020;26:1341-1343.ArticlePubMedPMC
Citations
Citations to this article as recorded by

- Reproduction Factor Based Latent Epidemic Model Inference: A Data-Driven Approach Using COVID-19 Datasets
Sujin Ahn, Minhae Kwon
IEEE Journal of Biomedical and Health Informatics.2023; 27(3): 1259. CrossRef - 코로나19 핵심 지표 산출체계 국제 비교 및 활용도 제고 방안 연구
나애 이, 연경 김, 승필 정, 우주 이, 주환 오, 승식 황
Public Health Weekly Report.2023; 16(29): 973. CrossRef - The Impacts of Compact City Characteristics on COVID-19 Spreading Force : Focused on the Seoul Metropolitan Area
Haejun Hyun, Myungje Woo
Journal of Korea Planning Association.2023; 58(7): 5. CrossRef - COVID-19 early-alert signals using human behavior alternative data
Anasse Bari, Aashish Khubchandani, Junzhang Wang, Matthias Heymann, Megan Coffee
Social Network Analysis and Mining.2021;[Epub] CrossRef
 , Yeon-Kyung Kim2
, Yeon-Kyung Kim2 , Woo-Sik Son3
, Woo-Sik Son3 , Jong-Hoon Kim4
, Jong-Hoon Kim4 , Jungsoon Choi5
, Jungsoon Choi5 , Baeg-Ju Na6
, Baeg-Ju Na6 , Boyoung Park1
, Boyoung Park1 , Bo Youl Choi1
, Bo Youl Choi1


 KSE
KSE



 PubReader
PubReader ePub Link
ePub Link Cite
Cite